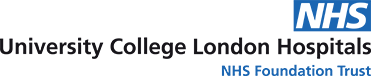This project is funded by NHSX and the National Institute for Health Research (NIHR) through the AI (artificial intelligence) in Health and Care Awards scheme, and is led by CRIU researchers Drs Anoop Shah and Wai Keong Wong.
Background
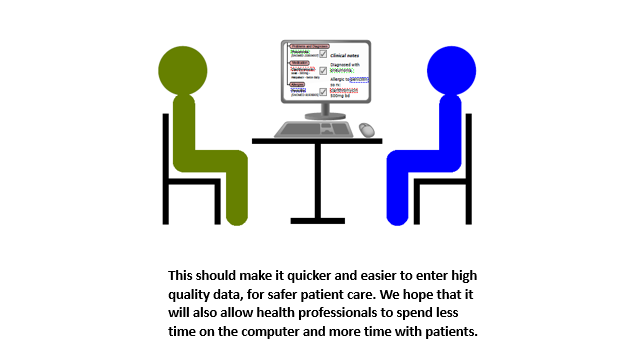
Data about people’s health stored in electronic health records (EHRs) can play an important role in improving the quality of patient care. Much of the information in EHRs is recorded in ordinary language without any restriction on format (“free text”), as this is the natural way in which people communicate. However, if this information were stored in a standardised, structured format, computers will also be able to process the information to help clinicians find and interpret information for better and safer decision making. This would enable EHR systems such as Epic, the system in place at UCLH since April 2019, to support clinical decision making. For instance, the system may be able to ensure that a patient is not prescribed a medicine they are allergic to.
The challenge
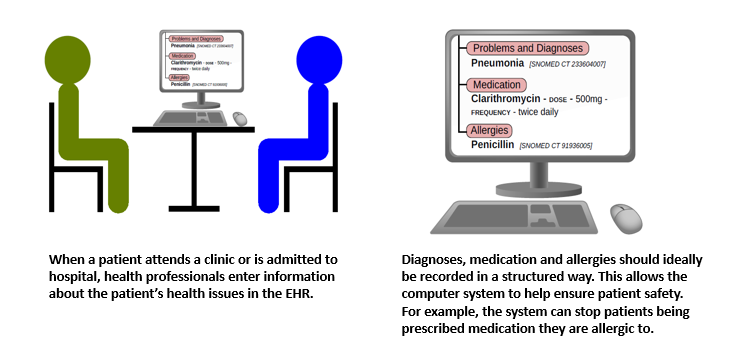
Free text may contain words and abbreviations which may be interpreted in more than one way, such as “HR”, which can mean “Hour” or “Heart Rate”. Free text may also contain negations; for example, a diagnosis may be mentioned in the text but the rest of the sentence might say that it was ruled out. Although computers can be used to interpret free text, they cannot always get it right, so clinicians will always have to check the results to ensure patient safety. Expressing information in a structured way can avoid this problem, but has a big disadvantage - it can be time-consuming for clinicians to enter the information. This can mean that information is incomplete, or clinicians are so busy on the computer that they do not have time to listen to their patients.
Meeting the need
The aim of MiADE is to develop a system to support automatic conversion of the clinician’s free text into a structured format. The clinician can check the structured data immediately, before making it a formal part of the patient’s record. The system will record a patient’s diagnoses, medications and allergies in a structured way, using NHS-endorsed clinical data standards (e.g. FIHR and SNOMED CT). It will use a technique called Natural Language Processing (NLP). NLP has been used by research teams to extract information from existing EHRs, but has rarely been used to improve the way information is entered in the first place. Our NLP system will continuously learn and improve as more text is analysed and checked by clinicians.
We will first test the system in University College London Hospitals, where a new EHR system called Epic is in place. We will study how effective it is, and how clinicians and patients find it when it is used in consultations. Based on feedback, we will make improvements and install it for testing at a second site (Great Ormond Street Hospital). Our aim is for the system to be eventually rolled out to more hospitals and doctors’ surgeries across the NHS.
The UCL/UCLH Clinical and Research Informatics Unit (CRIU)
MiADE will be facilitated by the CRIU, a collaboration between UCLH and the UCL Institute of Health Informatics. The CRIU team bridge the gap between university research and patient care, and aim to harness the potential of UCLH patient data for research to improve care. The CRIU will work closely with the Digital Research Environment at Great Ormond Street Hospital (DRIVE) on this project.
Immediate benefit for patient care
Better structured data in health records will have many advantages for safe, effective patient care. A clear, structured summary of diagnoses and treatments is invaluable for shared care, and when handing over the care of a patient (such as between shifts in hospital, or when patients are discharged or transferred between care settings).
Structured data can also enable EHR systems to assist clinical decision making. Many EHR systems include automatic warnings of medication allergies and interactions, and automated reminders for monitoring of chronic diseases. All these decision support aids rely on accurate, structured data to be present in the EHR. Clinical error is a major source of patient harm.
The proposed NLP system will enable the advantages of structured data to be realised while avoiding the disadvantage of the burden on clinicians entering the data. We will also study how the system affects the patient experience of consultations, which may hopefully improve if clinicians have to spend less time entering data into the computer.
Benefits for research to improve future care
Electronic health records are used in a large number of research studies for patient benefit. All these studies rely on high quality data; missing data can introduce bias and might result in inaccurate study outcomes which can lead to patient harm. If clinically-recorded data are not sufficiently complete, time-consuming retrospective data entry may be needed. For example, ongoing research projects on COVID-19 at UCLH are having to rely on retrospective manual data extraction for comorbidities and smoking status.
Clinical trials are vital for developing and evaluating new treatments, but many trials fail to recruit an adequate number of participants. Automated algorithms can help to detect patients eligible for certain trials, but only if the EHR contains high quality data. We believe that all patient groups will benefit, but sicker patients or those with more complex clinical histories may benefit more, as they may be at more risk of harm from clinical error due to missing information.
Benefits to the NHS and the wider population
MiADE will make it easier to use data for purposes beyond individual care. Although existing NLP approaches are being applied to health record databases, data need to be validated before they are used for decisions that may impact patients. Our approach enables immediate validation, and the data can be used for operational research, service planning, audit, safety monitoring, and clinical coding in near real time. Potential benefits include better care derived from better research, a reduction in resources needed for clinical coding, and more equitable allocation of resources.
We have estimated potential improvement in structured data entry from an audit of a recent data enhancement project, which found that during the COVID-19 pandemic, only two-thirds of diagnoses for patients admitted to UCLH were recorded in a structured way. Although the commercial sector is also interested in NLP solutions for healthcare, our publicly-funded NHS-led approach will ensure that all the intellectual property derived from this work, such as of the NLP models (developed using thousands of NHS patient records) remains within the NHS, and can benefit all future NHS patients. We will make the application code open source, and trained NLP models will be available for sharing with other NHS sites under appropriate data governance arrangements.
Throughout this project we are committed to maintaining the highest standards of data security in order to protect patient confidentiality.

Contact us
We welcome comments, enquiries and other contributions you might wish to send us. Please contact the team at uclh.miade@nhs.net