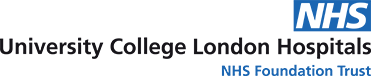Our mission is to ensure that data analytics of real-world clinical data generates evidence that is integrated with clinical practice and so delivers benefit to individual patients and their caregivers, leading to positive health outcomes.
We have a number of collaborations and partnerships with organisations worldwide and welcome the opportunity to work together to deliver the benefits of our research to patients and the public.
Current collaborations include:
This project is funded by NHSX and the National Institute for Health Research (NIHR) through the AI (artificial intelligence) in Health and Care Awards scheme, and is led by CRIU researchers Drs Anoop Shah and Wai Keong Wong.
Background
Data about people’s health stored in electronic health records (EHRs) can play an important role in improving the quality of patient care. Much of the information in EHRs is recorded in ordinary language without any restriction on format (“free text”), as this is the natural way in which people communicate. However, if this information were stored in a standardised, structured format, computers will also be able to process the information to help clinicians find and interpret information for better and safer decision making. This would enable EHR systems such as Epic, the system in place at UCLH since April 2019, to support clinical decision making. For instance, the system may be able to ensure that a patient is not prescribed a medicine they are allergic to.
The challenge
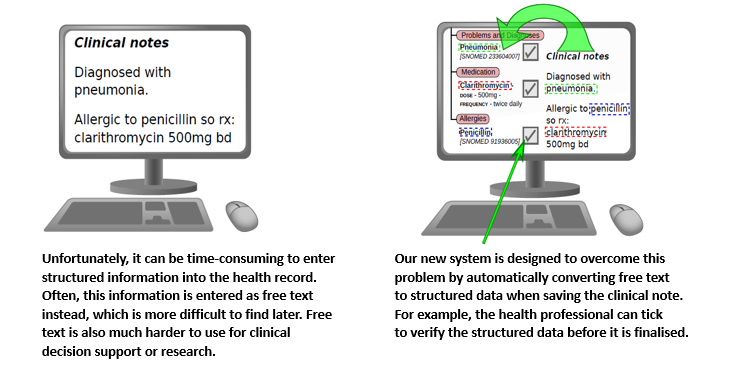
Free text may contain words and abbreviations which may be interpreted in more than one way, such as “HR”, which can mean “Hour” or “Heart Rate”. Free text may also contain negations; for example, a diagnosis may be mentioned in the text but the rest of the sentence might say that it was ruled out. Although computers can be used to interpret free text, they cannot always get it right, so clinicians will always have to check the results to ensure patient safety. Expressing information in a structured way can avoid this problem, but has a big disadvantage - it can be time-consuming for clinicians to enter the information. This can mean that information is incomplete, or clinicians are so busy on the computer that they do not have time to listen to their patients.
Meeting the need
The aim of MiADE is to develop a system to support automatic conversion of the clinician’s free text into a structured format. The clinician can check the structured data immediately, before making it a formal part of the patient’s record. The system will record a patient’s diagnoses, medications and allergies in a structured way, using NHS-endorsed clinical data standards (e.g. FIHR and SNOMED CT). It will use a technique called Natural Language Processing (NLP). NLP has been used by research teams to extract information from existing EHRs, but has rarely been used to improve the way information is entered in the first place. Our NLP system will continuously learn and improve as more text is analysed and checked by clinicians.
We will first test the system in University College London Hospitals, where a new EHR system called Epic is in place. We will study how effective it is, and how clinicians and patients find it when it is used in consultations. Based on feedback, we will make improvements and install it for testing at a second site (Great Ormond Street Hospital). Our aim is for the system to be eventually rolled out to more hospitals and doctors’ surgeries across the NHS.
The UCL/UCLH Clinical and Research Informatics Unit (CRIU)
MiADE will be facilitated by the CRIU, a collaboration between UCLH and the UCL Institute of Health Informatics. The CRIU team bridge the gap between university research and patient care, and aim to harness the potential of UCLH patient data for research to improve care. The CRIU will work closely with the Digital Research Environment at Great Ormond Street Hospital (DRIVE) on this project.
Immediate benefit for patient care
Better structured data in health records will have many advantages for safe, effective patient care. A clear, structured summary of diagnoses and treatments is invaluable for shared care, and when handing over the care of a patient (such as between shifts in hospital, or when patients are discharged or transferred between care settings).
Structured data can also enable EHR systems to assist clinical decision making. Many EHR systems include automatic warnings of medication allergies and interactions, and automated reminders for monitoring of chronic diseases. All these decision support aids rely on accurate, structured data to be present in the EHR. Clinical error is a major source of patient harm.
The proposed NLP system will enable the advantages of structured data to be realised while avoiding the disadvantage of the burden on clinicians entering the data. We will also study how the system affects the patient experience of consultations, which may hopefully improve if clinicians have to spend less time entering data into the computer.
Benefits for research to improve future care
Electronic health records are used in a large number of research studies for patient benefit. All these studies rely on high quality data; missing data can introduce bias and might result in inaccurate study outcomes which can lead to patient harm. If clinically-recorded data are not sufficiently complete, time-consuming retrospective data entry may be needed. For example, ongoing research projects on COVID-19 at UCLH are having to rely on retrospective manual data extraction for comorbidities and smoking status.
Clinical trials are vital for developing and evaluating new treatments, but many trials fail to recruit an adequate number of participants. Automated algorithms can help to detect patients eligible for certain trials, but only if the EHR contains high quality data. We believe that all patient groups will benefit, but sicker patients or those with more complex clinical histories may benefit more, as they may be at more risk of harm from clinical error due to missing information.
Benefits to the NHS and the wider population
MiADE will make it easier to use data for purposes beyond individual care. Although existing NLP approaches are being applied to health record databases, data need to be validated before they are used for decisions that may impact patients. Our approach enables immediate validation, and the data can be used for operational research, service planning, audit, safety monitoring, and clinical coding in near real time. Potential benefits include better care derived from better research, a reduction in resources needed for clinical coding, and more equitable allocation of resources.
We have estimated potential improvement in structured data entry from an audit of a recent data enhancement project, which found that during the COVID-19 pandemic, only two-thirds of diagnoses for patients admitted to UCLH were recorded in a structured way. Although the commercial sector is also interested in NLP solutions for healthcare, our publicly-funded NHS-led approach will ensure that all the intellectual property derived from this work, such as of the NLP models (developed using thousands of NHS patient records) remains within the NHS, and can benefit all future NHS patients. We will make the application code open source, and trained NLP models will be available for sharing with other NHS sites under appropriate data governance arrangements.
Throughout this project we are committed to maintaining the highest standards of data security in order to protect patient confidentiality.

New paper published on data gaps in electronic health record (EHR) systems
CRIU researchers Anoop Shah and Leilei Zhu, and medical student Jordan Poulos are authors of a new paper just published in The International Journal of Medical Informatics. The aim of the study was to conduct an audit to evaluate the completeness of diagnosis recording in 'problem lists' in a hospital electronic health record (EHR) system during the COVID-19 pandemic.
The problem list is a feature of EHR, which allows diagnoses and other clinical information to be stored in a structured way, with the aim to facilitate handovers and continuity of care, thus supporting clinical decision-making and enabling research.
The main findings of this study concluded that one year after implementation of a comprehensive EHR system at UCLH, the recording of medical history on the structured problem list for inpatients was incomplete, with almost 40% of important diagnoses mentioned only in the free text notes. This figure is comparable to studies carried out at other sites and internationally.
UCLH has begun a project funded by the NIHR to develop natural language processing technology to convert diagnoses entered in free text into coded terms in real time, which will appear in problem lists.
The researchers also identified other ways to close the ‘data gap’ – including through additional training in structured data. The researchers discuss potential reasons for their findings – suggesting that recording diagnoses in free text may have become the established way of working with a new EHR at UCLH, which was difficult to shift later.
Read the open access paper: Poulos J, Zhu L, Shah AD. Data gaps in electronic health record (EHR) systems: An audit of problem list completeness during the COVID-19 pandemic. International Journal of Medical Informatics 2021; 150(104452). https://doi.org/10.1016/j.ijmedinf.2021.104452
New paper looking at the interplay between heart disease and Covid-19
People with heart failure and atrial fibrillation who are hospitalised with Covid-19 are at greater risk of death, according to new findings from CAPACITY-COVID. The paper, published online on medRxiv.org, suggests that heart failure increases the risk of death by around 43%, while atrial fibrillation increases the risk by around 14%.
People with severe heart failure had the highest mortality rate amongst patients with heart disease, but interestingly other heart conditions, including having had a previous myocardial infarction (heart attack), were not related to an increased risk of death – showing that different types of heart disease affect Covid-19 risk differently.
Prof Bryan Williams, Chair of Medicine at University College London (UCL) and Director of Research at UCL Hospitals, who led the UK contribution to the study said: “This study provides important detail on the risk from Covid in people with pre-existing heart diseases and shows that for most heart conditions there is no increased risk of death in people hospitalised with Covid but for those with more severe heart failure or to a lesser extent, those with atrial fibrillation, the risk of death is increased. This helps refine our understanding of particular at risk populations for preventive strategies such as priority vaccination programmes.”
Prof Folkert Asselbergs, Consultant Cardiologist at University Medical Center Utrecht, is leading the European study overall.
Researchers monitored the disease progression of more than 10,000 patients hospitalised with Covid-19 in 16 countries across Europe between March and November 2020. They compared data from patients with and without heart disease to gain more insight into the role of these diseases in Covid-19 patients. Patients were followed from hospital admission to discharge.
The study, which was designated a National Institute for Health Research/British Heart Foundation Covid flagship research programme, also found:
- Mortality risk increased with age, both in patients with heart disease and those without
- There were no differences seen in Covid-19 symptoms (e.g. fever, cough, shortness of breath) between patients with heart disease and those without heart disease
- Complaints of smell loss – known to be a symptom of Covid-19 – occurred mainly in younger patients and much less in patients over 65
Prof Folkert Asselbergs, who is also Director of the UCLH BRC Clinical and Research Informatics Unit, said: “A key finding was that different types of heart disease affect mortality risk differently – so we shouldn’t necessarily talk about people with heart disease as one homogenous group when it comes to discussing risk.”
A key follow up piece of work will look at the long term effects of Covid-19 on the heart. To study this, Covid-19 patients who have been in hospital for Covid-19 will have ultrasound scans over time, and will also be surveyed.
This will shed light on whether there is an increased risk of heart disease in the years after hospitalisation and how often long-term complaints such as palpitations and chest pain occur.
Prof Asselbergs said: “This is a crucial area to explore as we go forward, because Covid-19 is still a new disease, and we will want to keep an eye on what impact it might have on people in 5, 10 or 20 years’ time – or even longer.”
Data requests for COVID-19-related initiatives
The Data Explorer (DEX) is a new platform developed by the Clinical and Research Informatics Unit (CRIU) to formally request access to health data which a UCLH member of staff (including honorary members) would not normally have access to.
During the pandemic peaks, we put in place a streamlined process for COVID-19-related initiatives, to rapidly review and support information needs. The requester was asked to complete a one-page form, including details of the people involved and the required data points. Then, the now-defunct COVID-19 Data Access Committee (DAC) was tasked with reviewing each request and further liaise with the requester should any questions arise concerning their data needs.
For enquiries on data access specifically involving COVID-19-related work, please contact the Joint Research Office: uclh.randd@nhs.net.
For assistance with the use of DEX or other questions about the platform, please contact the DEX team: uclh.dex@nhs.net.
COVID-19 Data Access Committee (DAC)
This now-defunct UCLH committee was created in April 2020 to assess, authorise and prioritise all data access initiatives related to COVID-19 response. The DAC also acted as a sub-committee of the UCLH COVID-19 Research Strategy and Compassionate Treatment Group, and during the pandemic peaks used to meet twice-weekly.
HIC and COVID-19
Since the start of the coronavirus pandemic, the NIHR Health Informatics Collaborative (HIC) Cardiovascular theme has been leading on a data collaboration to allow cardiovascular questions to be answered in relation to COVID-19.
As part of the HIC, NHS trusts with BRCs have been collating and sharing routinely-collected data to support research. The HIC trusts have now started to collate and share data to support research into COVID-19. The NIHR HIC COVID-19 dataset includes routinely-collected data on: admissions; blood tests; virology; microbiology; prescribing; medicines administration; orders; vital signs; and critical care. The aim is to collect data for every patient with a COVID test.
The secondary databases will provide data to support collaborative, translational research, particularly in the area of cardiovascular medicine. The curated data will also support other key initiatives, including the CAPACITY-COVID UK registry and DECOVID.
CAPACITY-COVID: UK-wide collection of cardiovascular complications due to COVID-19 infection
This British Heart Foundation flagship project is led by Professor Bryan Williams, Director of the NIHR University College Hospitals Biomedical Research Centre (BRC), with strong regional support.
The Clinical and Research Informatics Unit (CRIU) provides the data hosting infrastructure by establishing a local REDCap instance and server and coordinating data collection and transfer from participating sites, including outputs from the ISARIC-WHO REDCap uploads where applicable.
REDCap, the Clinical Data Entry tool supporting the project, is hosted by AIMES Management Services Ltd.
Can COVID-19 cause acute heart problems?
The research team will use hospital data from across the UK to understand how cardiovascular disease and risk factors increase the risk of developing severe complications in patients affected by COVID-19. Researchers will also explore which acute cardiovascular complications are most common in patients affected by the disease.
This UK project links with the European-wide CAPACITY (Cardiac complicAtions in Patients with SARS Corona vIrus 2 regisTrY; chief investigator Professor Folkert Asselbergs UCL/UMC Utrecht), a collaborative consortium seeking to establish an international registry of patients with COVID-19 with the aim to answer questions on the role of cardiovascular disease in this pandemic. The register is an extension of the Case Record Form that was released by ISARIC (International Severe Acute Respiratory and Emerging Infection Consortium) and WHO (World Health Organization). Since the launch of the registry, 88 centres across 17 countries have registered to join CAPACITY.
International Severe Acute Respiratory and emerging Infection Consortium (ISARIC)
ISARIC is a global federation of clinical research networks whose main purpose is to prevent illness and deaths from infectious disease outbreaks. ISARIC aims to provide a coordinated research response to outbreak-prone infectious diseases.
The emergence of the novel coronavirus (SARS-CoV-2) in December 2019 and the resulting pandemic of a severe coronavirus disease, COVID-19, has led to many ISARIC member networks, who are involved in patient-based research, to generate an evidence-based response to COVID 19.
ISARIC 4C (Coronavirus Clinical Characterisation Consortium) is a UK-wide consortium of doctors and scientists committed to answering urgent questions about COVID-19 quickly, openly, and for the benefit of all. Funded by a grant from UKRI (MRC), ISARIV 4C aims to addresses questions such as:
- How long are people infectious, and what body fluids are infectious?
- What puts people at higher risk of severe illness?
- What is the best way to diagnose the disease?
- Who should we treat early with drugs, and which drugs cause harm?
- Does the immune system in some patients do more harm than good?
- What other infections(such as pneumonia or flu) happen at the same time?
ISARIC 4C is funded to obtain serial samples from 300 cases and single samples for a further 1000. A full list of samples can be found in the UK clinical characterisation protocol. This is a generic, sleeping protocol that is designed to give fast responses with global harmonisation during outbreaks.
ISARIC 4C will share samples to get answers as fast as possible. The intention is to use every drop of every sample to have the biggest possible impact on the COVID-19 crisis. Any investigators with the ability to contribute can access ISARIC 4C data and samples. The ISARIC 4C study will provide a foundation for other studies, such as clinical trials of new treatments, to help better understand the best way to use interventions.
ISARIC 4C is one the UK-wide NIHR Urgent Public Health Priority studies for hospitalised patients with COVID-19.
This work uses data provided by patients and collected by the National Health Services (NHS) of the United Kingdom as part of their care and support #DataSavesLives.
At UCLH, the CRIU team is providing expertise and front-end build in Epic to automate streamline and improve the quality of the data captured directly from the EHR system.
Members of our team have contributed and provided lead authors to a range of high-impact publications...
- Ahmad T, Freeman JV, Asselbergs FW. Can advanced analytics fix modern medicine's problem of uncertainty, imprecision, and inaccuracy? Eur J Heart Fail. 2019 Jan;21(1):86-89. doi: 10.1002/ejhf.1370. Epub 2018 Dec 10.
- Akyea, R. K., Leonardi-Bee, J., Asselbergs, F. W., Patel, R. S., Durrington, P., Wierzbicki, A. S., . . . Weng, S. F. Predicting major adverse cardiovascular events for secondary prevention: protocol for a systematic review and meta-analysis of risk prediction models. BMJ open 2020; 10(7), e034564. doi:10.1136/bmjopen-2019-034564.
- Al-Rubaish, A. M., Al-Muhanna, F. A., Alshehri, A. M., Al-Mansori, M. A., Alali, R. A., Khalil, R. M., . . . Al-Ali, A. K. Bedside testing of CYP2C19 gene for treatment of patients with PCI with antiplatelet therapy. BMC cardiovascular disorders 2020; 20(1), 268. doi:10.1186/s12872-020-01558-2.
- Angermann, C. E., Assmus, B., Anker, S. D., Asselbergs, F. W., Brachmann, J., Brett, M. E., . . . Böhm, M. Pulmonary artery pressure-guided therapy in ambulatory patients with symptomatic heart failure: the CardioMEMS European Monitoring Study for Heart Failure (MEMS-HF). European Journal of Heart Failure 2020; doi:10.1002/ejhf.1943.
- Asselbergs FW, Meijboom FJ. Big data analytics in adult congenital heart disease: why coding matters. Eur Heart J. 2019 Apr 1;40(13):1078-1080. doi: 10.1093/eurheartj/ehz089.
- Bagheri, A., Sammani, A., van der Heijden, P. G. M., Asselbergs, F. W., & Oberski, D. L. (2020). ETM: Enrichment by topic modeling for automated clinical sentence classification to detect patients' disease history. JOURNAL OF INTELLIGENT INFORMATION SYSTEMS, 21 pages. doi:10.1007/s10844-020-00605-w.
- Bean, Daniel M., et al. Network analysis of patient flow in two UK acute care hospitals identifies key sub-networks for A&E performance. PLOS ONE 12(10): e0185912. doi: 10.1371/journal.pone.0185912
- Bosman, L. P., Cadrin-Tourigny, J., Bourfiss, M., Aliyari Ghasabeh, M., Sharma, A., Tichnell, C., . . . Te Riele, A. S. J. M. Diagnosing arrhythmogenic right ventricular cardiomyopathy by 2010 Task Force Criteria: clinical performance and simplified practical implementation. Europace : European pacing, arrhythmias, and cardiac electrophysiology : journal of the working groups on cardiac pacing, arrhythmias, and cardiac cellular electrophysiology of the European Society of Cardiology 2020; 22(5), 787-796. doi:10.1093/europace/euaa039.
- de Boer, R. A., Nijenkamp, L. L. A. M., Silljé, H. H. W., Eijgenraam, T. R., Parbhudayal, R., van Driel, B., . . . van der Velden, J. Strength of patient cohorts and biobanks for cardiomyopathy research. Netherlands Heart Journal 2020; 28, 50-56. doi:10.1007/s12471-020-01456-4.
- Gho JMIH, Schmidt AF, Pasea L, Koudstaal S, Pujades-Rodriguez M, Denaxas S, Shah AD, Patel RS, Gale CP, Hoes AW, Cleland JG, Hemingway H, Asselbergs FW. An electronic health records cohort study on heart failure following myocardial infarction in England: incidence and predictors. BMJ Open. 2018 Mar 3;8(3):e018331. doi: 10.1136/bmjopen-2017-018331. Erratum in: BMJ Open. 2018 Mar 22;8(3):e018331corr1.
- Gorrell, Genevieve, et al. Bio-YODIE: A named entity linking system for biomedical text. arXiv preprint arXiv 2018; 1811.04860. https://arxiv.org/pdf/1811.04860.pdf.
- Groenhof TKJ, Asselbergs FW, Groenwold RHH, Grobbee DE, Visseren FLJ, Bots ML; UCC-SMART study group. The effect of computerized decision support systems on cardiovascular risk factors: a systematic review and meta-analysis. BMC Med Inform Decis Mak. 2019 Jun 10;19(1):108. doi: 10.1186/s12911-019-0824-x.
- Groenhof TKJ, Koers LR, Blasse E, de Groot M, Grobbee DE, Bots ML, Asselbergs FW, Lely AT, Haitjema S; UPOD; UCC-CVRM Study Groups. Data mining information from electronic health records produced high yield and accuracy for current smoking status. J Clin Epidemiol. 2020 Feb;118:100-106. doi: 10.1016/j.jclinepi.2019.11.006. Epub 2019 Nov 12.
- Groenhof TKJ, Kofink D, Bots ML, Nathoe HM, Hoefer IE, Van Solinge WW, Lely AT, Asselbergs FW, Haitjema S. Low-Density Lipoprotein Cholesterol Target Attainment in Patients With Established Cardiovascular Disease: Analysis of Routine Care Data. JMIR Med Inform. 2020 Apr 2;8(4):e16400. doi: 10.2196/16400.
- Hagenbeek, F. A., Pool, R., van Dongen, J., Draisma, H. M., Hottenga, J. J., Willemsen, G., . . . Boomsma, D. I. Heritability estimates for 361 blood metabolites across 40 genome-wide association studies (vol 11, 39, 2020). NATURE COMMUNICATIONS 2020; 11(1), 1 page. doi:10.1038/s41467-020-15276-y
- Harris S, Shi S, Brealey D, MacCallum NS, Denaxas S, Perez-Suarez D et al. Critical Care Health Informatics Collaborative (CCHIC): Data, tools and methods for reproducible research: A multi-centre UK intensive care database. Int J Med Inform 2018; 112: 82–9. https://doi.org/10.1016/j.ijmedinf.2018.01.006.
- Heidemann, B. E., Koopal, C., Bots, M. L., Asselbergs, F. W., Westerink, J., & Visseren, F. L. J. (2020). The relation between VLDL-cholesterol and risk of cardiovascular events in patients with manifest cardiovascular disease. International Journal of Cardiology. doi:10.1016/j.ijcard.2020.08.030.
- Helgadottir, A., Thorleifsson, G., Alexandersson, K. F., Tragante, V., Thorsteinsdottir, M., Eiriksson, F. F., . . . Stefansson, K. Genetic variability in the absorption of dietary sterols affects the risk of coronary artery disease. Eur Heart J. 2020; 41(28), 2618-2628. doi:10.1093/eurheartj/ehaa531.
- Hemingway H, Asselbergs FW, Danesh J, Dobson R, Maniadakis N, Maggioni A, van Thiel GJM, Cronin M, Brobert G, Vardas P, Anker SD, Grobbee DE, Denaxas S; Innovative Medicines Initiative 2nd programme, Big Data for Better Outcomes, BigData@Heart Consortium of 20 academic and industry partners including ESC. Big data from electronic health records for early and late translational cardiovascular research: challenges and potential. Eur Heart J. 2018 Apr 21;39(16):1481-1495. doi: 10.1093/eurheartj/ehx487.
- Jackson, Richard, et al. CogStack-experiences of deploying integrated information retrieval and extraction services in a large National Health Service Foundation Trust hospital. BMC medical informatics and decision making 2018; 18(47). doi: 10.1186/s12911-018-0623-9.
- Kaura A, Arnold AD, Panoulas V, Glampson B, Davies J, Mulla A, Woods K, Omigie J, Shah AD, Channon KM, Weber JN, Thursz MR, Elliott P, Hemingway H, Williams B, Asselbergs FW, O'Sullivan M, Lord GM, Melikian N, Lefroy DC, Francis DP, Shah AM, Kharbanda R, Perera D, Patel RS, Mayet J. Prognostic significance of troponin level in 3121 patients presenting with atrial fibrillation (The NIHR Health Informatics Collaborative TROP-AF study). J Am Heart Assoc. 2020 Apr 7;9(7):e013684. doi: 10.1161/JAHA.119.013684. Epub 2020 Mar 26.
- Kaura A, Panoulas V, Glampson B, Davies J, Mulla A, Woods K, Omigie J, Shah AD, Channon KM, Weber JN, Thursz MR, Elliott P, Hemingway H, Williams B, Asselbergs F, O'Sullivan M, Kharbanda R, Lord GM, Melikian N, Patel RS, Perera D, Shah AM, Francis DP, Mayet J. Association of troponin level and age with mortality in 250 000 patients: cohort study across five UK acute care centres. BMJ. 2019 Nov 20;367:l6055. doi: 10.1136/bmj.l6055.
- Kaura A, Sterne JAC, Trickey A, Abbott S, Mulla A, Glampson B, Panoulas V, Davies J, Woods K, Omigie J, Shah AD, Channon KM, Weber JN, Thursz MR, Elliott P, Hemingway H, Williams B, Asselbergs FW, O'Sullivan M, Lord GM, Melikian N, Johnson T, Francis DP, Shah AM, Perera D, Kharbanda R, Patel RS, Mayet J. Invasive versus non-invasive management of older patients with non-ST elevation myocardial infarction (SENIOR-NSTEMI): a cohort study based on routine clinical data. Lancet. 2020 Aug 29;396(10251):623-634. doi: 10.1016/S0140-6736(20)30930-2.
- Klooster, C. C. V. T., Bhatt, D. L., Steg, P. G., Massaro, J. M., Dorresteijn, J. A. N., Westerink, J., . . . Visseren, F. L. J. Predicting 10-year risk of recurrent cardiovascular events andcardiovascular interventions in patients with established cardiovascular disease: results from UCC-SMART and REACH. International Journal of Cardiology 2020; doi:10.1016/j.ijcard.2020.09.053.
- Koudstaal S, Pujades-Rodriguez M, Denaxas S, Gho JMIH, Shah AD, Yu N, Patel RS, Gale CP, Hoes AW, Cleland JG, Asselbergs FW, Hemingway H. Prognostic burden of heart failure recorded in primary care, acute hospital admissions, or both: a population-based linked electronic health record cohort study in 2.1 million people. Eur J Heart Fail. 2017 Sep;19(9):1119-1127. doi: 10.1002/ejhf.709. Epub 2016 Dec 23.
- Kraljevica Z, Searleaf T, Shek A Roguski L,Noor K, Bean et al.Multi-domain Clinical Natural Language Processing with MedCAT: the Medical Concept Annotation Toolkit.
Artificial Intelligence in Medicine. Available online 1 May 2021. https://doi.org/10.1016/j.artmed.2021.102083. - Linschoten M, Asselbergs FW. CAPACITY-COVID: a European registry to determine the role of cardiovascular disease in the COVID-19 pandemic. Eur Heart J. 2020 Apr 8:ehaa280. doi: 10.1093/eurheartj/ehaa280. Epub ahead of print.
- Linschoten M, Asselbergs FW, CAPACITY-COVID collaborative consortium, LEOSS Study Group. Clinical presentation, disease course and outcome of COVID-19 in hospitalized patients with and without pre-existing cardiac disease – a cohort study across sixteen countries. doi: https://doi.org/10.1101/2021.03.11.21253106 (pre-print server).
- Linschoten, M., Kamphuis, J. A. M., & Asselbergs, F. W. (2020). Cardiovascular adverse events following treatment for non-Hodgkin lymphoma reply. Lancet Haemotology, 7(8), E557-E558.
- Lopez-Sainz, A., Dominguez, F., Lopes, L. R., Ochoa, J. P., Barriales-Villa, R., Climent, V., . . Dooijes, D. Clinical Features and Natural History of PRKAG2 Variant Cardiac Glycogenosis. Journal of the American College of Cardiology 2020; 76(2), 186-197. doi:10.1016/j.jacc.2020.05.029
- Mahmoodi, B. K., Tragante, V., Kleber, M. E., Holmes, M. V., Schmidt, A. F., McCubrey, R. O., . . . Patel, R. Association of Factor V Leiden with Subsequent Atherothrombotic Events: A GENIUS-CHD Study of Individual Participant Data. Circulation 2020; doi:10.1161/CIRCULATIONAHA.119.045526.http://10.1161/CIRCULATIONAHA.119.045526
- Meiring C, Dixit A, Harris S, MacCallum NS, Brealey DA, Watkinson PJ, Jones A et al. Optimal intensive care outcome prediction over time using machine learning. PLoS One 2018; 13(11):e0206862. doi: 10.1371/journal.pone.0206862. eCollection 2018.
- Noor K, Roguski L, Bai X, Handy A, Klapaukh R, Folarin A, Romao L, Matteson J, Lea N, Zhu L, Asselbergs F, Wong W, Shah A, Dobson R; Deployment of a Free-Text Analytics Platform at a UK National Health Service Research Hospital: CogStack at University College London Hospitals JMIR Med Inform 2022;10(8):e38122; URL: https://medinform.jmir.org/2022/8/e38122 DOI: 10.2196/38122
- Oskarsson, G. R., Oddsson, A., Magnusson, M. K., Kristjansson, R. P., Halldorsson, G. H., Ferkingstad, E., . . . Stefansson, K. Predicted loss and gain of function mutations in ACO1 are associated with erythropoiesis. COMMUNICATIONS BIOLOGY 2020; 3(1). doi:10.1038/s42003-020-0921-5.
- Palmer E, Post B, Klapaukh R, Marra G, Harris S, MacCallum NS et al. The association between supraphysiologic arterial oxygen levels and mortality in critically ill patients. A multicenter observational cohort study. Am J Respir Crit Care Med 2019; 200(11):1373–80, https://www.atsjournals.org/doi/10.1164/rccm.201904-0849OC
- Pei, J., Harakalova, M., Treibel, T. A., Lumbers, R. T., Boukens, B. J., Efimov, I. R., . . . Cheng, C. H3K27ac acetylome signatures reveal the epigenomic reorganization in remodeled non-failing human hearts.. Clin Epigenetics 2020; 12(1), 106. doi:10.1186/s13148-020-00895-5.
- Pool, R., Hagenbeek, F. A., Hendriks, A. M., van Dongen, J., Willemsen, G., de Geus, E., . . . Slagboom, P. E. Genetics and Not Shared Environment Explains Familial Resemblance in Adult Metabolomics Data. TWIN RESEARCH AND HUMAN GENETICS 2020; 23(3), 145-155. doi:10.1017/thg.2020.53.
- Poulos J, Zhu L, Shah AD. Data gaps in electronic health record (EHR) systems: An audit of problem list completeness during the COVID-19 pandemic. International Journal of Medical Informatics 2021; 150(104452). https://doi.org/10.1016/j.ijmedinf.2021.104452.
- Sammani, A., Kayvanpour, E., Bosman, L. P., Sedaghat-Hamedani, F., Proctor, T., Gi, W. -T., . . . Asselbergs, F. W. Predicting sustained ventricular arrhythmias in dilated cardiomyopathy: a meta-analysis and systematic review. ESC HEART FAILURE 2020; doi:10.1002/ehf2.12689.
- Savarese, G., Settergren, C., Schrage, B., Thorvaldsen, T., Löfman, I., Sartipy, U., . . . Lund, L. H. Comorbidities and cause-specific outcomes in heart failure across the ejection fraction spectrum: A blueprint for clinical trial design. International Journal of Cardiology 2020; doi:10.1016/j.ijcard.2020.04.068.
- Schmidt, A. F., Finan, C., Gordillo-Marañón, M., Asselbergs, F. W., Freitag, D. F., Patel, R. S., . . . Hingorani, A. D. Genetic drug target validation using Mendelian randomisation. Nature Communications 2020; 11(1). doi:10.1038/s41467-020-16969-0.
- Searle, Tom, et al. MedCATTrainer: A biomedical free text annotation Interface with active learning and research use case specific customisation. Proceedings of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP): System Demonstrations 2019; D19-3024. doi: 10.18653/v1/D19-3024.
- Shi S, Pérez-Suárez D, Harris S, MacCallum N, Brealey D, Singer M et al. Critical care data processing tools. The Journal of Open Source Software 2017; 2(20):513. doi: 10.21105/joss.00513.
- Smith DA, Wang T, Oliver Freeman O, Charles Crichton C, Hizni Salih H, Philippa Clare Matthews PC, Jim Davies J, Kinga Anna Várnai KA, Shaw TA, Drumright LN, Romão L, David Ramlakan D, Higgins F, Weir A, Nastouli E, Agarwa K, Gelson W, Cooke GS, Barnes E. National Institute for Health Research Health Informatics Collaborative: development of a pipeline to collate electronic clinical data for viral hepatitis research. BMJ Health & Care Informatics 2020;27:e100145. doi: 10.1136/bmjhci-2020-100145.
- Stolfo D, Uijl A, Benson L, Schrage Budim M, Asselbergs FW, Koudstaal S, Sinagra G, Dahlström U, Rosano G, Savarese G. Association between beta-blocker use and mortality/morbidity in older patients with heart failure with reduced ejection fraction. A propensity score-matched analysis from the Swedish Heart Failure Registry. Eur J Heart Fail. 2020 Jan;22(1):103-112. doi: 10.1002/ejhf.1615. Epub 2019 Oct 23.
- Timmerman, N., de Kleijn, D. P. V., de Borst, G. J., den Ruijter, H. M., Asselbergs, F. W., Pasterkamp, G., . . . van der Laan, S. W. Family history and polygenic risk of cardiovascular disease: Independent factors associated with secondary cardiovascular events in patients undergoing carotid endarterectomy. Atherosclerosis 2020; doi:10.1016/j.atherosclerosis.2020.04.013.
- Tissot HC, Shah AD, Brealey D, Harris S, Agbakoba R, Folarin A, Romao L, Roguski L, Dobson R, Asselbergs FW. Natural language processing for mimicking clinical trial recruitment in critical care: a semi-automated simulation based on the LeoPARDS trial. IEEE J Biomed Health Inform. 2020; 24(10):2950-9. doi: 10.1109/JBHI.2020.2977925.
- Uijl A, Koudstaal S, Direk K, Denaxas S, Groenwold RHH, Banerjee A, Hoes AW, Hemingway H, Asselbergs FW. Risk factors for incident heart failure in age- and sex-specific strata: a population-based cohort using linked electronic health records. Eur J Heart Fail. 2019 Oct;21(10):1197-1206. doi: 10.1002/ejhf.1350. Epub 2019 Jan 7.
- Uijl, A., Lund, L. H., Vaartjes, I., Brugts, J. J., Linssen, G. C., Asselbergs, F. W., . . . Savarese, G. A registry-based algorithm to predict ejection fraction in patients with heart failure. ESC HEART FAILURE 2020, 10 pages. doi:10.1002/ehf2.12779.
- Van Den Berg, V. J., Umans, V. A. W. M., Brankovic, M., Oemrawsingh, R. M., Asselbergs, F. W., Van Der Harst, P., . . . Akkerhuis, K. M. Stabilization patterns and variability of hs-CRP, NT-proBNP and ST2 during 1 year after acute coronary syndrome admission: Results of the BIOMArCS study. Clinical Chemistry and Laboratory Medicine 2020;. doi:10.1515/cclm-2019-1320.
- van ’t Klooster, C. C., van der Graaf, Y., Ridker, P. M., Westerink, J., Hjortnaes, J., Sluijs, I., . . . Visseren, F. L. J. The relation between healthy lifestyle changes and decrease in systemic inflammation in patients with stable cardiovascular disease. Atherosclerosis 2020; 301, 37-43. doi:10.1016/j.atherosclerosis.2020.03.022.
- Why Unstructured Data Holds the Key to Intelligent Healthcare Systems. HIT Consultant. 2015. URL: https://hitconsultant.net/2015/03/31/tapping-unstructured-data-healthcares-biggest-hurdle-realized/ [accessed 2022-07-08]
- Willeit, P., Tschiderer, L., Allara, E., Reuber, K., Seekircher, L., Gao, L., . . . Lorenz, M. W. Carotid Intima-Media Thickness Progression as Surrogate Marker for Cardiovascular Risk: Meta-Analysis of 119 Clinical Trials Involving 100 667 Patients. Circulation 2020; 142(7), 621-642. doi:10.1161/CIRCULATIONAHA.120.046361
- Wu, Honghan, et al. SemEHR: A general-purpose semantic search system to surface semantic data from clinical notes for tailored care, trial recruitment, and clinical research. Journal of the American Medical Informatics Association 2018; 25(5):530–53. doi: 10.1093/jamia/ocx160.
- Wu, Honghan, et al. SemEHR: Surfacing semantic data from clinical notes in electronic health records for tailored care, trial recruitment, and clinical research. The Lancet 2017; 390(S97). doi: 10.1016/S0140-6736(17)33032-5.
At CRIU we work with and support a number of postgraduate students whose main research focus is clinical informatics, artificial intelligence and machine learning.
Reham AlDakhil
Project title:
“Exploring the factors that influence the clinical decision making when using electronic prescribing as part of the electronic health record (EHR)"
About my research:
Electronic prescribing (eP) systems have been advocated as a strategy and solution to reducing medication errors and improving patient safety. The evidence for the effectiveness of eP shows mixed outcomes, with some errors being reduced, newer computer related errors being introduced and emergence of various workarounds and unintended consequences on workflow. The degree to which errors may be reduced is often dependent on the level of sophistication of the system in providing clinical decision support. Basic clinical decision support systems (CDSS) involve a series of checks such as drug-allergy, drug-drug; drug-disease and drug-food interactions or simple dose checking using an underlying clinical database and are often presented in the form of interruptive alerts. Advanced clinical decision support may include complex dosing support for renal insufficiency and at extremes of age, guidance for medication-related laboratory testing, drug–pregnancy checking and other guided treatment decisions using order sets or pathways including use of algorithms.
Few studies have conducted robust assessments of factors that influence clinical decision making at point of care when using electronic prescribing systems as part of an integrated electronic health record system (EHRS).
The evidence for the effectiveness of electronic prescribing (eP) shows mixed outcomes; emerges of various workarounds, new types of computer-related errors being introduced and unintended consequences on workflow, with some errors being reduced. The involvement of various forms of decision support within the clinical decision support systems (CDSS) from the basic checks of the drug-drug interactions to the advance form of guided treatment decision using order sets including the use of algorithms.
Few studies have conducted robust assessments of factors that influence clinical decision making at the point of care when using electronic prescribing systems as part of an integrated electronic health record system (EHRS).
The PhD project will conduct a mixed-method approach to map the current evidence of the influence of these factors and will explore the potential of eP within the EHR systems to enhance the appropriate use of the system in the medication management process.
About me:
I obtained a BSc in Pharmaceutical Sciences from King Saud University, Riyadh. I worked as a hospital pharmacist at NGHA and was part of some automation projects with the ISD. I was awarded an MSc in Health Informatics, KSAU-HS, in 2018.
My main research interests are electronic prescribing, human factors and clinical decision making related to EHR.
Supervisors:
Professor Folkert Asselbergs
Dr Yogini Jani
Dr Wai Keong Wong
Dr Yang Chen
Project title:
“Learning Health Systems and decision support within acute cardiovascular care”
About my research:
My project aims to generate new insights into acute cardiovascular disease (CVD) management through analysis of electronic health record (EHR) data, existing structured datasets and the design and delivery of a pragmatic randomised clinical trial (RCT) embedded within the EHR.
For the trial, my specific focus will be in the area of fluid overload and heart failure. I plan to examine whether a clinician-facing clinical decision support system (CDSS) embedded in the EHR can modify behaviour and deliver a standardised intervention of fluid restriction compared to standard of care. Along with my supervisors and the trial team, we will measure the effects of our intervention on relevant clinical outcomes during an initial pilot phase and then determine the best strategy for taking the work forward.
About me:
I graduated in medicine from the University of Oxford in 2013. In 2017 I started my specialist training in Cardiology and was appointed as an NIHR academic clinical fellow at UCL. Since then, I have completed an Executive MSc in Health Economics, Outcomes and Management in Cardiovascular Sciences at the London School of Economics (LSE). My professional interest is in how decision-making occurs in both a clinical and research context, and the boundary between routine care, pragmatic trials within EHRs and traditional RCTs.
Supervisors:
Dr Tom Lumbers
Dr Anoop Shah
Prof Folkert Asselbergs
Katarzyna Dziopa
Project title:
"Predicting cardiovascular disease in type 2 diabetes patients utilizing contemporary electronic healthcare records”
About my research:
Cardiovascular disease is one of the major complications of T2DM, with 32.2% patients experiencing a CVD event annually. Despite the considerable CVD risk associated with T2DM, individual patients’ risk may differ markedly, between 1% to 28%. This individual variation suggests there is an (unmet) need for risk-stratified management, similar to CVD prevention in the general population. In this research we aim to explore the possibilities for CVD risk-stratification inT2DM patients by:
- Externally validating 20+ frequently used CVD prediction rules in T2DM patients.
- Using machine learning to identify relevant CVD features from high-dimensional data such as the UK biobank.
- Evaluating the utility of ensemble learning methodologies compared to existing prediction algorithms.
About me:
I have an MSc in Computer Science.
Supervisors:
Professor Folkert W. Asselbergs
Dr Floriaan Schmidt
Professor Nishi Chaturvedi
Joseph Farrington
Project title:
“AI-enabled blood transfusion system”.
About my research:
Blood products are central to many aspects of modern medical care. Ordering and supply of both red cell and platelet units are complicated by the requirement to use specific blood-group or platelet-group types. This is currently carried out by local experts using crude estimates, which leads to over or under ordering of specific blood groups to constitute stock. Patient care may suffer if blood products are not available and treatment is delayed or cancelled if products are not available.
In this project, we will explore various AI methods to build models of making accurate predictions of the blood product usage by learning from local experts and using actual data from NHS and UCLH. Based on the prediction model, optimal approaches of blood products ordering system can be developed and implemented in the long run. Specifically, integration of hospital and laboratory data can define a practical model that can help to order and reduce wastage, and it would be much more powerful if real-time data and deep learning techniques for prediction of demand are utilized. Furthermore, the AI models of blood products recommendation will be evaluated prospectively on the actual retrospective use at UCLH. The application of this technology at scale, would reduce blood wastage and reduce inappropriate use of universal donor blood. We believe that this project fits with the goals of the CDT as it uses applies AI to manage what is a scarce resource (donations from voluntary donors) to match against the transfusion need for patients. This is a unique collaboration between the national blood service (NHSBT), an NHS trust (UCLH) with a fully integrated comprehensive EHR (Epic) and a maturing research platform (EMAP).
About me:
I was awarded an MSc in Machine Learning, UCL, and a BA in Natural Sciences, University of Cambridge.
Supervisors:
Dr Ken Li, Machine Learning supervisor
Dr Wai Keong, Clinical Informatics supervisor
Matt Wilson
Project title:
“Exploiting natural clinical variation using observational and experimental methods to create an embedded learning healthcare system for Critical Care”
About my research:
Many routine clinical decisions in critical care lack a clear evidence base. In these situations of equipoise, clinicians base decisions on experience and acumen. As clinicians vary, so do decisions, contributing to variation in outcomes for patients.
Current practice is to reduce all variation via the application of guidelines, protocols and audit. However, this fails to preserve scenarios where clinicians deviate from guidelines justifiably, resulting in improved outcomes for patients.
My PhD will explore the feasibility of addressing this question using a combination of observational techniques to map existing variation and a novel 'nudge' tool as part of a randomised, embedded clinical trial of routine critical care interventions.
About me:
I graduated from St Bart's and The Royal London Medical School in 2011 and work as an anaesthetic registrar. I completed an MSc in Health Data Science in 2018 and am currently an MRC Doctoral Training Partnership student. My main research interests include investigating causal relationships using natural experimental methods with observational data and advancing electronic health record systems for clinician learning and feedback.
Supervisors:
Professor Folkert Asselbergs
Dr Steve Harris
Dr Roma Klapaukh
Workforce Training and Capacity Building
We are strongly committed to enabling Life Learning and we support opportunities for learning outside the framework of a formal programme of study.
Clinician coders
A programme provided by the UCL Academic Careers Office (ACO) to support clinical academics and healthcare professionals to effectively use data science to complement their existing research. The programme consists of a series of two-day workshops where participants with little or no previous experience can do hands-on data science and computer programming with support from experts and their peers. The programme also foster multidisciplinary working and will help you to separate yourself from the pack in your next fellowship application. Workshops are announced on the Clinician Coders webpage and the ACO newsletter. Registration is required and a limited number of places is available on each workshop. To find out more, visit: https://www.ucl.ac.uk/school-life-medical-sciences/about-slms/office-vice-provost-health/academic-careers-office/career-schemes/clinician-coders
UCL Institute of Health Informatics short courses
The Institute runs a series of one- or two-day short courses each Spring, with the main focus on developing skills to fully harness the potential of electronic health records and translational data science. To find out more, visit: https://www.ucl.ac.uk/health-informatics/study/continuing-professional-development